Projects
Protein AFM Analysis and Lennard–Jones Potential Fitting
Quantitative force–distance analysis of protein AFM data using Lennard–Jones (LJ) potential modeling
Overview
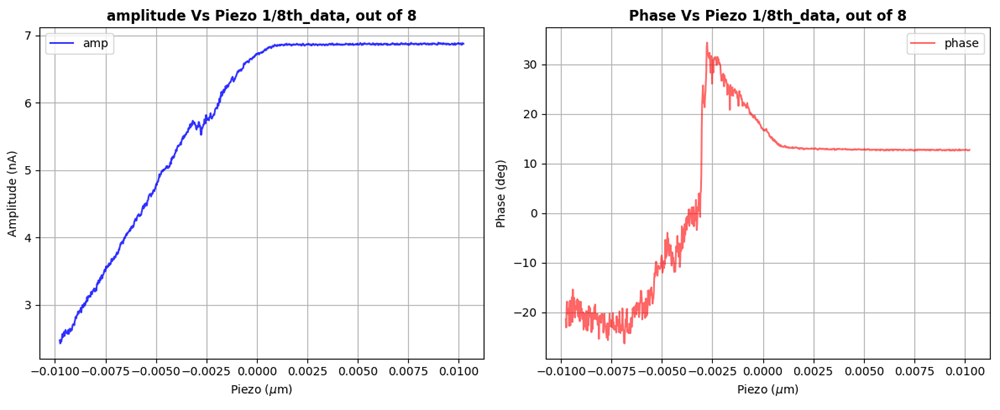
This project focuses on quantitative analysis of Atomic Force Microscopy (AFM) force–distance curves acquired from protein samples.
The goal is to extract meaningful interaction parameters by fitting experimental data to the Lennard–Jones (LJ) potential, a standard model for intermolecular forces.
The workflow combines AFM data preprocessing, numerical modeling, and nonlinear curve fitting to study nanoscale protein–surface interactions.
Problem Statement
AFM force–distance measurements often contain:
- noise and instrumental artifacts,
- non-linear interaction regimes,
- difficulty in extracting physical parameters directly.
Interpreting these curves in a physically meaningful way requires:
- a robust preprocessing pipeline,
- a suitable interaction model,
- stable parameter estimation.
Approach
The analysis pipeline:
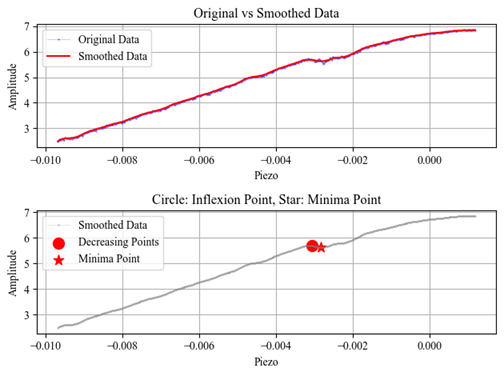
- Imports and preprocesses AFM force–distance data
- Identifies relevant interaction regions
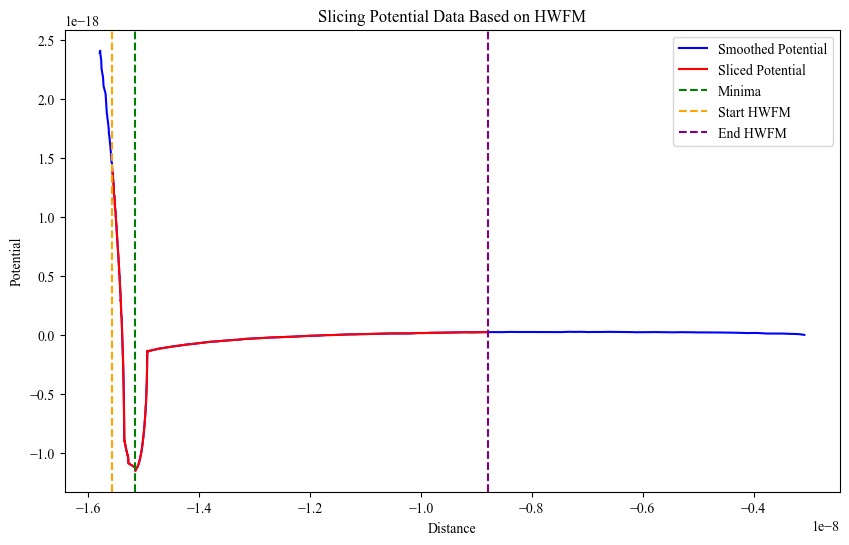
- Models tip–sample interaction using the Lennard–Jones potential
- Performs curve fitting to estimate interaction parameters
- Visualizes experimental vs fitted curves for validation
The fitting enables estimation of:
- equilibrium distance,
- interaction strength,
- repulsive and attractive force regimes.
Mathematical Model
The Lennard–Jones potential used in this project is given by:
$$
U(r) = 4\epsilon \left[\left(\frac{\sigma}{r}\right)^{12} - \left(\frac{\sigma}{r}\right)^6\right]
$$
Where:
- $r$ is the tip–sample separation
- $\epsilon$ represents interaction strength
- $\sigma$ is the equilibrium distance parameter
Force curves are derived from the gradient of the potential and fitted to experimental AFM data.
Technologies Used
- Python
- NumPy
- SciPy (optimization + curve fitting)
- Matplotlib (visualization)
- Jupyter Notebook
Results
- Successful fitting of AFM force–distance curves using an LJ model
- Clear separation of attractive vs repulsive interaction regimes
- Physically interpretable parameters extracted from experimental data
How to Run
- Clone the repository:
bash git clone https://github.com/mohdrafik/Protein-AFM-Analysis-and-Lennard-Jones-potential-fitting
Practical setup and Results