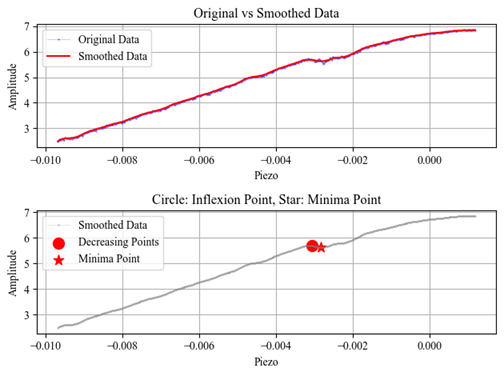
Protein AFM Analysis and Lennard–Jones Potential Fitting
Quantitative force–distance analysis of protein AFM data using Lennard–Jones (LJ) potential modeling
Project Summary:
- Solved the problem of extracting reliable physical parameters from noisy AFM force–distance measurements by designing an end-to-end Python data analysis and modeling pipeline.
- Built scalable scientific computing workflows using Python, NumPy, SciPy, and Matplotlib for preprocessing, feature extraction, nonlinear optimization, and model validation.
- Applied physics-informed modeling with the Lennard–Jones potential to estimate interaction strength and equilibrium distance in protein–surface interactions.
- Implemented numerical optimization, curve fitting, and statistical validation techniques to ensure robustness, reproducibility, and data quality.
- Delivered clear analytical visualizations and model–data comparisons to support data-driven decision making and experimental interpretation.
- Demonstrates strong expertise in data analysis, scientific Python, computational modeling, algorithm development, and translating experimental data into actionable insights for R&D and applied research environments.